Expert Review - (2021) Volume 0, Issue 0
Genetic variability of pedunculate oak (Quercus robur L.) at the Mediterranean margin of the distribution range
M. Memiševic Hodžic1, B. Fussi2, D. Gömöry3 and D. Ballian1,4*Abstract
In Bosnia and Herzegovina, pedunculate oak (Quercus robur L.) occurs at the southern margin of its distribution range, close to the glacial refugia of this species. To assess the patterns of genetic diversity distribution at the rear edge of the Holocene colonization, we studied genetic variation in 20 pedunculate oak populations using 14 allozyme loci. Despite considerable differences among populations, neither the numbers of alleles nor genetic diversity showed any geographical trend within the studied area, although small isolated populations showed generally lower allelic richness. The Bayesian analysis of population structure indicated a kind of geographical pattern. We identified no signs of a recent bottleneck. The proximity to multiple glacial refugia explains the outcomes.
Keywords
Pedunculate oak, allozyme, genetic variability, marginal populations, rear edge.
Introduction
Pedunculate oak (Quercus robur L.) is an essential component of forests in Bosnia and Herzegovina (BiH) (Begovic, 1960; Memiševic, 2008). However, current occurrences covering around 10,000 ha represent only a fraction of once-massive forest complexes, removed or turned into coppices by uncontrolled logging (Begovic 1960). In BiH, pedunculate oak is found mainly in mixed forests with hornbeam; pure stands occur only on specific sites (Stefanovic et al., 1983). The distribution range is continuous only in the northern part of riverain forests along the Sava river, although more significant occurrences can also be found in the other river valleys. Towards the Adriatic Sea, the increasing influence of Mediterranean climate and the mountainous character of the landscape make the range progressively fragmented, local populations become smaller, sometimes even individual trees are dispersed over the landscape (Memiševic 2008). However, even in the karstic regions of southwestern BiH, isolated larger populations can be found, e.g., at the Glasinac plateau near Sokolac or a marsh forest near Livno (Ballian et al., 2010). There is, however, no clear boundary between compact riparian oakwoods and fragmented sub-Mediterranean occurrences; the transition is relatively smooth.
The pedunculate oak in BiH is situated at the periphery of the range. Peripheral populations situated close to glacial refugia (the so-called ‘rear edge’ of the Holocene colonization; Hampe & Petit, 2005) are often specific in terms of gene pools. In zoochorous species such as oaks, allelic richness is typically the highest in refugial areas, where populations have persisted throughout the Pleistocene (not necessarily without shifts at a local or regional scale) and accumulated mutations. Allelic richness usually gradually declines towards the front edge of colonization due to recurrent founder events (Comps et al. 2001). Consequently, populations at southern range margins tend to contain unique genetic resources, even though not all preserved alleles need to have adaptive significance (Kelleher et al., 2015); this broader spectrum of gene variants may allow a species to persist under environmental fluctuations. Channel & Lomolino (2000) concluded that most species currently occur in the periphery of their historical-geographical ranges. In terms of gene conservation, this conclusion underlines the importance of southern-edge populations under the ongoing global climate change (Forest Europe, 2020).
Therefore, very few molecular-marker studies in pedunculate oak have covered BiH. Ballian et al. (2010) studied the variability of remnant populations and tree groups of pedunculate oak in BiHemploying nuclear microsatellites despite a small territory under study, and the use of nuclear markers reflecting pollen dispersal, solid genetic differentiation, and variation in allelic richness was recorded. Another study relied on seed-dispersed chloroplast genes: Slade et al. (2008) analyzed chloroplast DNA variation in four oak species (Q. robur, Q. petraea, Q. pubescens, and Q. frainetto) in a broader area of the central Balkans using PCR-RFLP markers. They detected the highest haplotype diversity in the Neretva river valley and suggested several refugia in the territory of BiH. Such a suggestion is not unrealistic. For understandable reasons, the most extensive study of chloroplast DNA variation in oaks accomplished within the FAIROAK project during the late 1990s included only three Bosnian populations from the Sava plain. However, although the southern and central Balkans was largely uncovered, the study concluded that primary refugia of oaks were present in this area (Bordács et al., 2002; Petit et al., 2002a, b).
Isolation and small population sizes resulting in solid genetic drift in refugial populations have led to differentiation at sometimes very small geographical scales. This differentiation is still discernible in maternally inherited portions of genomes, especially in zoochorous species and in mountainous landscapes (Papageorgiou et al., 2014). Naturally, gene flow by pollen gradually wipes out such geographical patterns in paternally or biparentally inherited genes. This study used the example of pedunculate oak in BiH as a region proximate to glacial refugia of oaks to detect the traces of such ancient differentiation and to provide valuable information for the regulation of the production and use of forest reproductive materials and conservation in situ and ex situ of pedunculate oak in BiH.
Materials and Methods
The study relies on a provenance experiment with pedunculate oak in BiH. Seeds were collected in autumn 2007 in twenty natural populations with a minimum of 10 trees per population (Table 1) and sown in spring 2009. In February 2014, twigs with dormant buds were collected from 50 plants per provenance and subjected to allozyme analysis.
| Provenance | Abbreviation | Latitude | Longitude | Altitude | Populationsize† |
|---|---|---|---|---|---|
| Bijeljina | Bije | 44º 43' 50'' | 19º 13' 30'' | 93 | L |
| BosanskaDubica | BDub | 45º 06' 24'' | 16º 40' 32'' | 145 | L |
| BosanskaGradiška | BGra | 45º 07' 04'' | 17º 19' 03'' | 91 | L |
| BosanskiBrod | BBro | 45º 05' 27'' | 18º 00' 38'' | 84 | L |
| BosanskoGrahovo | BGrh | 44º 01' 05'' | 16º 38' 24'' | 703 | S |
| Bugojno | Bugo | 44º 06' 00'' | 17º 26' 31'' | 537 | S |
| Drvar | Drva | 44º 23' 39'' | 16º 21' 54'' | 462 | S |
| HrgoviSrebrenik | HSre | 44º 49' 06'' | 18º 34' 11'' | 133 | L |
| Jelah | Jela | 44º 39' 09'' | 17º 56' 46'' | 181 | L |
| Kaćuni | Kacu | 44º 03' 59'' | 17º 56' 13'' | 443 | M |
| Ključ | Klju | 44º 30' 56'' | 16º 48' 42'' | 260 | S |
| Kotor Varoš | KVar | 44º 39' 07'' | 17º 21' 35'' | 252 | M |
| Miljevina | Milj | 43º 31' 06'' | 18º 38' 56'' | 627 | S |
| Mrkonjić Grad | MGra | 44º 27' 04'' | 16º 58' 42'' | 753 | S |
| Mutnica | Mutn | 44º 58' 55'' | 15º 50' 54'' | 270 | M |
| Olovo | Olov | 44º 07' 44'' | 18º 36' 11'' | 542 | S |
| Sokolac | Soko | 43º 55' 17'' | 18º 48' 53'' | 866 | S |
| Stojčevac | Stoj | 43º 48' 40'' | 18º 17' 25'' | 506 | S |
| Žepče | Zepc | 44º 25' 35'' | 18º 03' 10'' | 224 | M |
| Živinice | Zivi | 44º 27' 58'' | 18º 41' 09'' | 216 | L |
Table 1. Analyzed pedunculate oak provenances.
Enzymes were extracted from dormant buds and separated using starch-gel electrophoresis. Ten isoenzyme systems were used (E.C. numbers and controlled loci given in parentheses): alanine aminopeptidase (E.C. 3.4.11.1.; Aap-A), aspartate aminotransferase (E.C. 2.6.1.1., Aat-B), fluorescent a-esterase (E.C. 3.1.1.1.; Fest-A, Fest-C), alcohol dehydrogenase (E.C. 1.1.1.1; Adh-A), isocitrate dehydrogenase (E.C. 1.1.1.42.; Idh-A, Idh-B, Idh-C), menadione reductase (E.C. 1.6.1.99.2.; Mnr-A), 6-phosphogluconate dehydrogenase (E.C. 1.1.1.44.; 6-pgdh-A, 6-pgdh-B), phosphoglucose isomerase (E.C. 5.3.1.9.;Pgi-B), phosphoglucomutase (E.C. 2.7.5.1.; Pgm-B), and shikimate dehydrogenase (E.C. 1.1.1.25.; Skdh-A). Electrophoretic and staining procedures followed Konnert et al. (2004).
POPGENE 1.32 (Yeh et al., 1997) was used to calculate the parameters of intrapopulation variation. As the variation of sample sizes among populations was minimal, a total number of alleles was used to characterize allelic richness. Fixation indices and Weir & Cockerham’s unbiased estimator of the coefficient of genetic differentiation (Weir & Cockerham, 1984) were calculated using GENEPOP 4.3 (Raymond & Rousset, 1995). Deviations from Hardy-Weinberg equilibrium (HWE) were tested using exact tests (Guo & Thompson, 1992) and combined across loci using the Fisher’s method; again, calculations were carried out using GENEPOP 4.3.
To determine a genetic bottleneck, BOTTLENECK 1.2.02 was used (Cornuet & Luikart, 1996). The expected heterozygosity distribution was obtained by simulating the coalescent process both under the infinite allele model (IAM) and the two-phase model (TPM) with the proportion of stepwise mutations of 50%. Typically, IAM is suitable for allozyme loci. However, quite regular intervals between the relative migration rates of protein fractions produced by different isozyme alleles indicate stepwise changes in the net charge (Richardson et al. 1986); in fact, the stepwise mutation model (SMM) was initially developed for protein variants detectable by electrophoresis, differing in electric charge (Ohta & Kimura, 1973). Therefore, we found the application of TPM, which is a mixture of IAM and SMM, justified. The significance of heterozygote surplus or deficiency on the expectation under mutation-drift equilibrium was tested by the sign and Wilcoxon tests.
Bayesian analysis of population structure was conducted using STRUCTURE 2.3.4 (Pritchard et al. 2000; Hubisz et al. 2009), ten times for each K=1 to 10, with a burn-in period of 100,000 subsequent 500,000 iterations to determine the number of clusters. The admixture model was used, along with sampling location as prior information (LOCPRIOR), to assist the clustering. The optimal number of groups was determined based on the posterior probabilities of cluster membership and the ?K measure (Evanno et al., 2005) using the STRUCTURE HARVESTER program (Earl & von Holdt, 2012). Clusters were aligned using CLUMPP 1.1.2 (Jakobsson & Rosenberg, 2007) using the Greedy algorithm with 10,000 random permutations.
Allozyme loci represent coding sequences expressed in functional proteins (enzymes), so they may be subject to selection. To identify the signals of selection, the FST-outlier approach was applied. Bayescan v. 2.1 (Foll & Gaggiotti, 2008) was used for the computations, with 20 pilot runs of 5000 iterations each and a burn-in of 50,000 iterations followed by 50,000 iterations to estimate the posterior distribution with a thinning interval of 10. The evidence of selection relied on Bayes factors, measuring odds for selection model versus neutral model derived from posterior probabilities of each of the models. Prior odds for the neutral model were set to 10 (default). Posterior odds of more than 32:1 in favor of the selection model (i.e., log10(odds)>1.5) are to be considered strong evidence for selection. False discovery rate (FDR) was controlled through the q-value, which measures the minimum FDR at which this locus may become significant (the expected proportion of false positives among outliers is defined in the context of multiple testing across all polymorphic loci), while the strength and direction of selection (diversifying vs. balancing) was quantified by the α coefficient (Foll&Gaggiotti 2008).
Isolation by distance was tested according to Rousset (1997). As recommended for a two-dimensional case, regression of the estimates of FST/(1–FST) against the logarithm of distance was evaluated using GENEPOP v.4.3 (Raymond & Rousset, 1995). As the colonization trails were unknown, geographical coordinates were converted to orthogonal UTM coordinates, and simple air distances (Euclidean) between populations were used.
Results
The number of populations within each size category is too small to test the effects of population size on genetic variation levels explicitly. Variation ranges are significant for all parameters; for instance, the coefficient of variation among populations is almost 11% for expected heterozygosity. In spite of this, certain trends are visible. Level of polymorphism as measured by the proportion of polymorphic loci or mean number of alleles per locus was the highest in large populations (mostly riparian forests along the Sava and other rivers in the northern part of BiH), while the medium-sized and small isolated populations (located mostly in more mountainous regions) show less polymorphism (Table 2). This is only partly true for genetic diversity. Populations are mostly at HWE; in four cases (riparian Bos. Dubica, small Bos. Grahovo, Drvar, Miljevina) heterozygote deficiency was observed, while Olovo (small) exhibited heterozygote excess. We found also a marginally significant heterozygote excess in Stojcevac (small) (Table 2). Despite interpopulation variation, none of the parameters shows a clear geographical trend or pattern.
| Population | N | PP | na | ne | He | FIS | P |
|---|---|---|---|---|---|---|---|
| large populations | |||||||
| Bijeljina | 49.71 | 92.9 | 2.93 | 1.46 | 0.238 | 0.081 | 0.8964 |
| Bosanska Dubica | 49.14 | 85.7 | 2.36 | 1.36 | 0.203 | 0.225 | <0.0001 |
| Bosanska Gradiška | 48.86 | 92.9 | 2.79 | 1.38 | 0.211 | 0.035 | 0.9880 |
| Bosanski Brod | 49.43 | 92.9 | 3.14 | 1.53 | 0.261 | 0.010 | 0.3348 |
| Hrgovi Srebrenik | 49.79 | 92.9 | 2.79 | 1.45 | 0.241 | 0.006 | 0.4799 |
| Jelah | 50.00 | 92.9 | 2.64 | 1.53 | 0.280 | -0.004 | 0.9161 |
| Živinice | 49.79 | 100.0 | 2.79 | 1.49 | 0.273 | 0.025 | 0.6018 |
| average ± S.D. | 92.9±4.1 | 2.78±0.24 | 1.46±0.07 | 0.244±0.030 | |||
| medium populations | |||||||
| Kaćuni | 49.71 | 100.0 | 2.79 | 1.50 | 0.257 | -0.009 | 0.5937 |
| Kotor Varoš | 49.86 | 78.6 | 2.36 | 1.39 | 0.229 | -0.033 | 0.5200 |
| Mutnica | 49.57 | 92.9 | 2.79 | 1.37 | 0.205 | 0.029 | 0.9574 |
| Žepče | 47.57 | 92.9 | 2.57 | 1.33 | 0.198 | -0.068 | 0.8900 |
| average ± S.D. | 91.1±9.0 | 2.62±0.21 | 1.40±0.07 | 0.222±0.027 | |||
| small populations | |||||||
| Bosansko Grahovo | 49.14 | 78.6 | 2.36 | 1.44 | 0.225 | 0.166 | <0.0001 |
| Bugojno | 48.64 | 85.7 | 2.43 | 1.46 | 0.232 | 0.020 | 0.9237 |
| Drvar | 50.00 | 92.9 | 3.07 | 1.50 | 0.261 | 0.072 | 0.0474 |
| Ključ | 49.64 | 100.0 | 2.86 | 1.51 | 0.248 | 0.043 | 0.1606 |
| Miljevina | 47.36 | 92.9 | 3.07 | 1.48 | 0.252 | 0.151 | 0.0025 |
| Mrkonjić Grad | 46.79 | 78.6 | 2.71 | 1.45 | 0.226 | 0.086 | 0.5111 |
| Olovo | 49.64 | 85.7 | 2.36 | 1.41 | 0.239 | -0.208 | 0.0276 |
| Sokolac | 48.93 | 100.00 | 2.79 | 1.48 | 0.255 | 0.033 | 0.2047 |
| Stojčevac | 49.93 | 78.6 | 2.36 | 1.35 | 0.187 | -0.163 | 0.0555 |
| average ± S.D. | 88.1±8.8 | 2.67±0.30 | 1.45±0.05 | 0.236±0.022 | |||
| overall average ± S.D. | 90.4±7.4 | 2.70±0.26 | 1.44±0.06 | 0.236±0.026 | |||
Table 2. Parameters of genetic variability for the 20 analyzed provenances.
Tests of a recent bottleneck have not identified a heterozygosity excess in any population (Table 3). In contrast, a few populations exhibited heterozygosity deficits (i.e., a recent population expansion). Strictly consistent were the outcomes of tests only in Bos. Gradiška, where both the sign test and the Wilcoxon test showed significant heterozygosity deficiency under both TPM and IAM. In the other three populations (Žepce, Drvar, and Kljuc) three tests were significant, and the fourth was marginally significant (P<0.10). There is, however, no association between the outcomes of the bottleneck tests and the current population size.
| Locus | P-value | log10(PO) | q | α |
|---|---|---|---|---|
| Aap-A | 1 | 1000 | 0 | -1.2720 |
| Aat-B | 0.9926 | 2.1299 | 0.0025 | -1.4143 |
| Adh-A | 0.9896 | 1.9776 | 0.0044 | -1.0767 |
| Fest-A | 0.1575 | -0.7283 | 0.2780 | -0.1122 |
| Fest-C | 0.5561 | 0.0979 | 0.1348 | -0.5565 |
| Idh-A | 1 | 1000 | 0 | -1.4168 |
| Idh-B | 0.0663 | -1.1490 | 0.3889 | -0.0289 |
| Idh-C | 0.8243 | 0.6714 | 0.0387 | -0.7097 |
| Mnr-A | 0.6941 | 0.3559 | 0.0832 | -0.5116 |
| 6pdg-A | 0.0550 | -1.2354 | 0.4743 | 0.0071 |
| 6pgd-B | 0.1573 | -0.7289 | 0.3345 | 0.1485 |
| Pgi-B | 0.0576 | -1.2138 | 0.4351 | -0.0135 |
| Pgm-B | 0.2839 | -0.4018 | 0.2074 | -0.1893 |
| Skdh-A | 0.0534 | -1.2486 | 0.5080 | 0.0073 |
Table 3. Identification of the signals of selection based on the Bayesian FST-outlier approach.
In the analysis under the STRUCTURE procedure, the ΔK measure (Evanno et al., 2005) indicated K=5 as the most probable number of groups. However, a secondary peak was observed at K=2 and a high ΔK was found also for K=3 (Fig. 1). However, there is no geographical or other interpretable pattern with 5 clusters. There are a few populations with the predominance of a particular cluster (Bos. Grahovo, Jelah, Kotor Varoš, Olovo), but the remaining populations exhibit a mixture of all gene pools, with no association with population size or geographical position (Fig. 2). The same applies to K = 2, the proportion of two gene pools is almost the same everywhere.
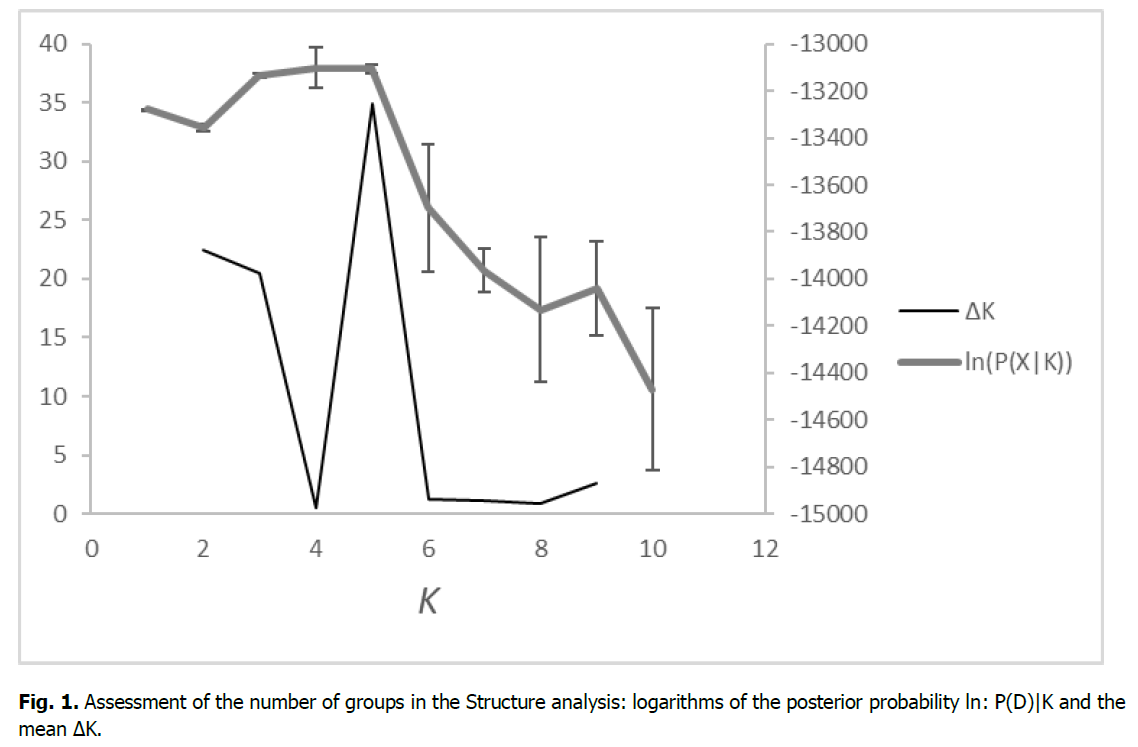
Figure 1: Assessment of the number of groups in the Structure analysis: logarithms of the posterior probability ln: P(D)|K and the mean ΔK.
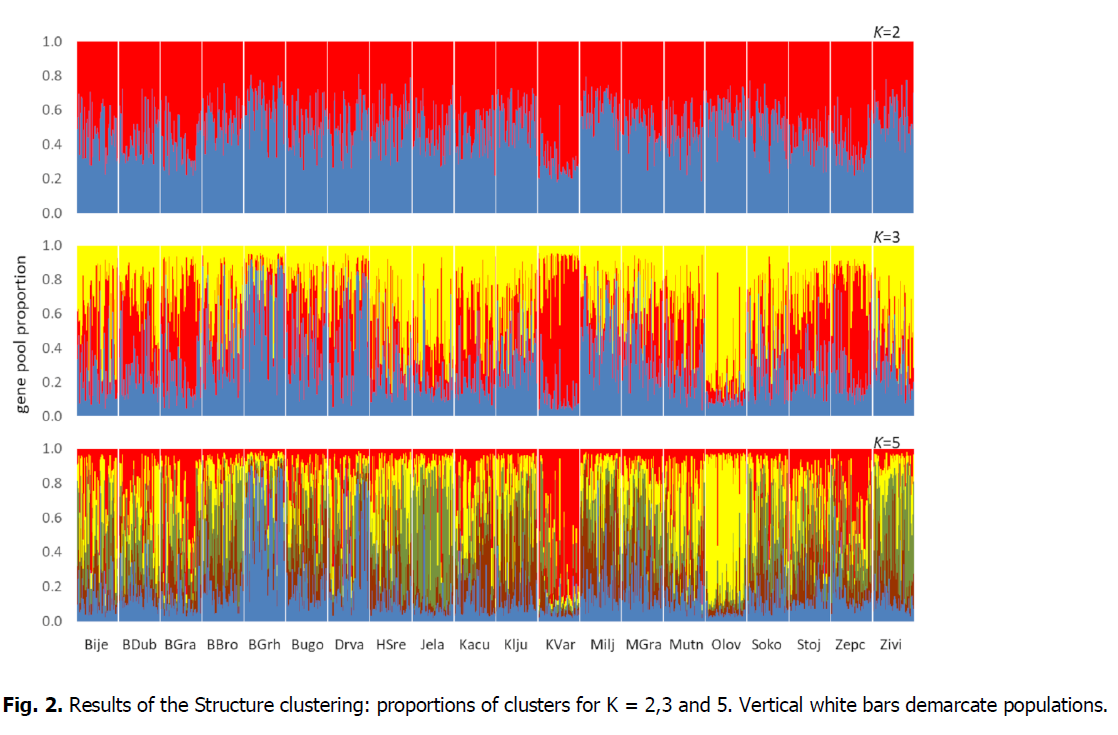
Figure 2: Results of the Structure clustering: proportions of clusters for K = 2,3 and 5. Vertical white bars demarcate populations.
A kind of pattern can be recognized at K=3, where contiguous groups of populations exhibit similar profiles. One gene pool predominates in the mountains of western BiH, another one in riparian populations along the Sava in the northwest, a third one again in the mountains north and Northeast of Sarajevo. Nevertheless, most local groups are interspersed by single populations exhibiting a deviating profile of gene pools. With the absence of any region-wide trend, it is not surprising that no isolation by distance was observed. The slope of the regression of pairwise FST on the logarithm of geographical distance is close to zero (b=-0.0035, 95%CI=[-0.0107, 0.0070]), and the Mantel test derived from 100,000 permutations was also non-significant (P =0.5251). However, the overall level of differentiation is quite high (FST=0.075).
BayeScan shows signals of selection at a few loci (Aap-A, Idh-A, Aat-B, Adh-A); the resulting posterior odds give ‘very strong’ to ‘decisive’ support for the model including selection (Foll & Gaggiotti, 2008), while the negative α-coefficients indicate balancing selection in all cases. However, considering the really small number of loci studied, these results need to be taken with caution (Table 4).
| Population | Sign test | Wilcoxon test | ||||
|---|---|---|---|---|---|---|
| IAM | TPM | H deficiency | H excess | |||
| IAM | TPM | IAM | TPM | |||
| Bijeljina | 0.1153 | 0.0946 | 0.0341 | 0.0164 | 0.9713 | 0.9867 |
| Bosanska Dubica | 0.2383 | 0.1983 | 0.1902 | 0.1018 | 0.8303 | 0.9119 |
| Bosanska Gradiška | 0.0483 | 0.0301 | 0.0471 | 0.0239 | 0.9598 | 0.9801 |
| Bosanski Brod | 0.1077 | 0.0850 | 0.0839 | 0.0402 | 0.9268 | 0.9659 |
| Bosansko Grahovo | 0.2816 | 0.2309 | 0.3823 | 0.1826 | 0.6499 | 0.8398 |
| Bugojno | 0.2365 | 0.0745 | 0.2593 | 0.1167 | 0.7651 | 0.8982 |
| Drvar | 0.0349 | 0.0207 | 0.0839 | 0.0164 | 0.9268 | 0.9867 |
| Hrgovi Srebrenik | 0.1371 | 0.0278 | 0.0732 | 0.0287 | 0.9364 | 0.9761 |
| Jelah | 0.5321 | 0.4488 | 0.3424 | 0.1879 | 0.6823 | 0.8302 |
| Kaćuni | 0.0277 | 0.0173 | 0.1479 | 0.0969 | 0.8662 | 0.9137 |
| Ključ | 0.0309 | 0.0201 | 0.0969 | 0.0393 | 0.9137 | 0.9662 |
| Kotor Varoš | 0.3194 | 0.2560 | 0.3501 | 0.1602 | 0.6812 | 0.8608 |
| Miljevina | 0.1012 | 0.0236 | 0.0636 | 0.0287 | 0.9451 | 0.9761 |
| Mrkonjić Grad | 0.2254 | 0.1861 | 0.0737 | 0.0415 | 0.9385 | 0.9663 |
| Mutnica | 0.1524 | 0.1197 | 0.0471 | 0.0199 | 0.9598 | 0.9836 |
| Olovo | 0.5476 | 0.4067 | 0.3386 | 0.3110 | 0.6890 | 0.7153 |
| Sokolac | 0.4099 | 0.0198 | 0.1083 | 0.0209 | 0.9031 | 0.9824 |
| Stojčevac | 0.1256 | 0.0894 | 0.1030 | 0.0415 | 0.9126 | 0.9663 |
| Žepče | 0.0548 | 0.0366 | 0.0341 | 0.0107 | 0.9713 | 0.9957 |
| Živinice | 0.2240 | 0.1660 | 0.2508 | 0.1206 | 0.7684 | 0.8917 |
Table 4. Tests of a recent bottleneck in the analyzed pedunculate oak populations.
Discussion
Direct comparisons of genetic variation parameters among studies are hardly feasible with a small and nonrandom sample of markers, which is the unavoidable fate of allozyme studies. Nevertheless, the comparison of allelic frequencies at identical loci indicates that allelic richness in Bosnian populations is slightly smaller than in other regions such as France (Streiff et al., 1998), Romania (Craciunesc et al., 2011; Curtu et al., 2011) or Slovakia (Gömöry et al., 2001), unless the populations are located at the very margin of the distribution range (Yakovlev, 2000; Vakkari et al., 2006; Redkina et al., 2008). However, the variation ranges of the number of alleles or the proportion of polymorphic loci are broad and apparently not correlated with population size. Even when, on average, the number of alleles in small isolated populations in mountainous regions is smaller than in large riparian populations, in individual cases (Drvar, Miljevina) allelic richness is high. Bottleneck tests confirm the absence of allele loss: whenever a deviation from the mutation-drift equilibrium was found, a deficiency, rather than an excess of heterozygosity, was observed, and this applies also to small populations. Apparently, isolation at the southwestern range margin is by far not perfect and gene flow at the local or regional level suffices to prevent genetic erosion.
Except for the population Bos. Dubica, all large and medium populations are at Hardy-Weinberg equilibrium, the fixation indices are near zero. Among small populations, the variation is bigger; although fixation indices are generally positive (significantly in three cases), heterozygote excess was also observed. In general, oaks (including pedunculate oak) are at HWE or exhibit slight heterozygote deficiency, whatever markers are used (Jensen et al., 2003; Belletti et al., 2005; Di Pietro et al., 2021).
Vakkari et al. (2006) observed heterozygote deficiency primarily in large stands, which they attributed to population substructure, spatial (continuous groups of relatives) or temporal (variation in flowering phenology). Later investigation (Vakkari et al., 2020) revealed strong spatial genetic structure in small populations at the northern range limit. The existence of family clusters caused by limited seed dispersal is the most plausible explanation of positive inbreeding coefficients in our case. Nevertheless, heterozygote excess was also observed, although not as often; the most probable cause is gene flow by pollen from neighbouring genetically differentiated fragments (Degen et al., 2021).
High differentiation typically accompanies fragmentation; this was confirmed also in BiH. Despite the small territory under study, the overall coefficient of differentiation was quite high; however, this is quite typical for marginal oak populations (Vakkari et al., 2020; Degen et al., 2021; Di Pietro et al., 2021). Typically, interpopulation variation among isolated populations is attributed to genetic drift associated with reduced population sizes, i.e., the bottleneck effect in older fragments, and the founder effect in newly established populations. Indeed, most populations in BiH (except the riparian oakwoods in the country's north) are small, but bottleneck tests did not confirm this hypothesis. Of course, a set of 14 loci may be too small to detect signals of a bottleneck, but wherever the test outcomes are significant or nearly significant, they indicate a deficit rather than excess of heterozygosity, which means population expansion rather than decline.
We suggest population history as a more plausible explanation for the observed differentiation levels and patterns. The Structure analysis did not reveal any clear geographical trend, but the spatial distribution of gene pools is not completely chaotic, as is typical when genetic drift is the sole factor determining genetic structure (a typical example is Piceaomorica in BiH; Ballian et al., 2006). Whatever the number of clusters is considered (except K=2), a certain pattern is the distribution of cluster proportions can be identified, which may reflect the origins from different refugial sources. As already mentioned, the territory of BiH, and the southern Balkans in general, have been insufficiently covered by the hitherto phylogeographic studies of Q. robur. However, the distribution of maternally inherited chloroplast lineages and fossil pollen studies point towards the southern Balkans as an important refugial area and a source of recolonization of the present range of white oaks (subg. Quercus), especially in central and eastern Europe (Tzedakis, 1994; Petit et al., 2002a). A high chloroplast haplotype diversity was observedboth directly in BiH and in a close vicinity (Slade et al., 2008), documenting th e presence of multiple refugiain the broad area of the southern and southwestern Balkans. Paleobotanical sites yielding long fossil records even indicate an almost permanent presence of deciduous oaks in this region during the Late Pleistocene (Willis, 1992; Tzedakis, 1994; Stefanova & Ammann, 2003; Urban & Fuchs, 2005). Of course, both types of evidence apply to white oaks not specifically Q. robur.Not all colonization sources suggested by Slade et al. (2008) must have contributed to the gene pools of current pedunculate oak populations in BiH, and the colonization sources need not have been limited to the suggested ones. However, the geographically structured sharing of maternally inherited genome segments among white oak taxa indicates that different taxa may have shared their source refugial populations and have become reconstituted through recurrent introgression and selection (Petit et al., 1997; Cannon & Petit, 2019). We observed a certain degree of spatial continuity in the distribution of Structure profiles, which may be a remnant of colonization. Mosaic-like character of the spatial pattern of structure clusters is not surprising; in fact, a similar mosaic can be observed in the distribution of chloroplast haplotypes, which is a consequence of long-distance founding events at the outset of the Holocene colonization (Petit et al., 1997). In contrast to chloroplast markers, the initial differentiation of pollen-dispersed nuclear genes becomes gradually eroded by gene flow. Potentially, the stabilizing selection indicated by the FST-outliertests might also have contributed to gene pool homogenization. However, despite relatively long time elapsed since the colonization, the spatial pattern remains distinguishable.
Funding
MMH, DB: P4-0107 I LIFE SYSTEMIC INGenRes Bridge project, European Cooperation for Science and Technology, COST Action FP1202 Strengthening conservation: a key issue for the adaptation of marginal/peripheral populations of forest trees to climate change in Europe (MaP-FGR).
BF: Trees4Future project of the EU 7th Framework Programme
DG: Slovak Grant Agency for Science VEGA 1/0029/20
Conflict of Interest
The authors are not aware of any conflict of interests.
Acknowledgements
We would like to thank the P4-0107 I LIFE SYSTEMIC INGenRes Bridge project, Bavarian Office for Forest Genetics (AWG) for providing the laboratory facilities for performing this study, and the AWG staff, in particular Dr. Monika Konnert, for introducing MMH into the allozyme analysis. MMH and DB were financed through European Cooperation for Science and Technology, COST Action FP1202 Strengthening conservation: a key issue for the adaptation of marginal/peripheral populations of forest trees to climate change in Europe (MaP-FGR), BFby the Trees4Future project of the EU 7th Framework Programme, DG by the Slovak Grant Agency for Science project VEGA 1/0029/20.
References
Ballian, D., Belleti, P., Ferrazzini, D., Bogunić, F., Kajba, D. (2010). Genetic variability of pedunculate oak (Quercus robur L.) in BiH. Periodicum Biologorum, 112:353-362.
Ballian, D., Longauer, R., Mikić, T., Paule, L., Kajba, D., Gömöry, D. (2006). Genetic structure of a rare European conifer, Serbian spruce (Picea omorika (Panč.) Purk.). Plant Systematics and Evolution, 260:53-63.
Begović, B. (1960). Strani kapital u šumskoj privredi Bosne i Hercegovine za vrijeme otomanske vladavine. 5: 1–248.
Belletti, P., Leonardi, S., Monteleone, I., Piouani, P. (2005). Allozyme Variation in Different Species of Deciduous Oaks From Northwestern Italy. Silvae Genetica, 54:9-16.
Bordács, S., Popescu, F., Slade, D., Csaikl, U.M., Lesur, I., Borovics, A., Petit, R.J. (2002). Chloroplast DNA variation of white oaks in northern Balkans and in the Carpathian Basin. Forest Ecology and Management, 156:197-209.
Cannon, C.H., Petit, R.J. (2020). The oak syngameon: more than the sum of its parts. New Phytologist, 226:978-983.
Channell, R., Lomolino, M.V. (2000). Dynamic biogeography and conservation of endangered species. Nature, 403:84-86.
Comps, B., Gömöry, D., Letouzey, J., Thiébaut, B., Petit, R.J. (2001). Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics, 157:389-397.
Cornuet, J.M., Luikart, G. (1996). Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics, 144:2001-2014.
Craciunesc, I., Ciocîrlan, E., Sofletea, N., Curtu, A.L. (2011). Genetic diversity of pedunculate oak (Quercus robur L.) in Prejmer natural reserve. Bulletin of the Transilvania University of Brasov. Forestry, Wood Industry, Agricultural Food Engineering. Series II, 4:15.
Curtu, A.L., Gailing, O., Leinemann, L., Finkeldey, R. (2007). Genetic variation and differentiation within a natural community of five oak species (Quercus spp.). Plant Biology, 9:116-126.
Degen, B., Yanbaev, Y.A., Ianbaev, R.Y., Bakhtina, S.Y., Gabitova, A.A., Tagirova, A.A. (2021). Genetic Diversity and Differentiation of Northern Populations of Pedunculate Oak Based on Analysis of New SNP Markers. Russian Journal of Genetics, 57:374-378.
Di Pietro, R., Conte, A.L., Di Marzio, P., Fortini, P., Farris, E., Gianguzzi, L., Gailing, O. (2021). Does the genetic diversity among pubescent white oaks in southern Italy, Sicily and Sardinia islands support the current taxonomic classification?. European Journal of Forest Research, 140:355-371.
Earl, D.A. (2012). Structure harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources, 4:359-361.
Evanno, G., Regnaut, S., Goudet, J. (2005). Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology, 14:2611-2620.
Foll, M., Gaggiotti, O. (2008). A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics, 180:977-993.
Forest Europe. (2020). Adaptation to Climate Change in Sustainable Forest Management in Europe. Zvolen:Liaison Unit Bratislava.
Gömöry, D., Yakovlev, I., Zhelev, P., Jedináková, J., Paule, L. (2001). Genetic differentiation of oak populations within the Quercus robur/Quercus petraea complex in Central and Eastern Europe. Heredity, 86:557-563.
Guo, S.W., Thompson, E.A. (1992). Performing the exact test of Hardy-Weinberg proportion for multiple alleles. Biometrics, pp:361-372.
Hampe, A., Petit, R.J. (2005). Conserving biodiversity under climate change: the rear edge matters. Ecology Letters, 8:461-467.
Hubisz, M.J., Falush, D., Stephens, M., Pritchard, J.K. (2009). Inferring weak population structure with the assistance of sample group information. Molecular ecology resources, 9:1322-1332.
Jakobsson, M., Rosenberg, N.A. (2007). CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics, 23:1801-1806.
Jensen, J.S., Olrik, D.C., Siegismund, H.R., Lowe, A.J. (2003). Population genetics and spatial autocorrelation in an unmanaged stand of Quercus petraea in Denmark. Scandinavian Journal of Forest Research, 18:295-304.
Kelleher, C.T., de Vries, S.M., Baliuckas, V., Bozzano, M., Frýdl, J., Goicoechea, P. G., Stoyanov, P.Z. (2015). Approaches to the Conservation of Forest Genetic Resources: In Europe in the Context of Climate Change. Bioversity International.
Konnert, M., Fromm, M., Wimmer, T. (2004). Anleitung für Isoenzymuntersuchungen bei Stieleiche (Quercus robur) und Traubeneiche (Quercus petraea). Bund-Länder-Arbeitsgruppe „Erhaltung forstlicher Genressourcen “, Selbstverlag.
Memišević, M. (2008). Eksploatacija kao razlog nestanka hrasta lužnjaka (Quercus robur, L.) u periodu od 1878. do 1914. godine u Bosni i Hercegovini. Naše šume, 12:39-40.
Ohta, T., Kimura, M. (1973). A model of mutation appropriate to estimate the number of electrophoretically detectable alleles in a finite population. Genetics Research, 22:201-204.
Papageorgiou, A.C., Tsiripidis, I., Mouratidis, T., Hatziskakis, S., Gailing, O., Eliades, N.G.H., Finkeldey, R. (2014). Complex fine-scale phylogeographical patterns in a putative refugial region for Fagus sylvatica (Fagaceae). Botanical Journal of the Linnean Society, 174:516-528.
Petit, R.J., Brewer, S., Bordács, S., Burg, K., Cheddadi, R., Coart, E., Kremer, A. (2002). Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. Forest Ecology and Management, 156:49-74.
Petit, R.J., Csaikl, U.M., Bordács, S., Burg, K., Coart, E., Cottrell, J., Kremer, A. (2002). Chloroplast DNA variation in European white oaks: phylogeography and patterns of diversity based on data from over 2600 populations. Forest Ecology and Management, 156:5-26.
Petit, R.J., Pineau, E., Demesure, B., Bacilieri, R., Ducousso, A., Kremer, A. (1997). Chloroplast DNA footprints of postglacial recolonization by oaks. Proceedings of the National Academy of Sciences, 94:9996-10001.
Pritchard, J.K., Stephens, M., Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics, 155:945-959.
Raymond, M. (1995). GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Journal of Heredity, 86:248-249.
Redkina, N.N., Mullagulov, R.Y., Yanbaev, Y.A., Degen, B. (2008). Fine spatial structure of allozyme genotypes in isolated population of pedunculate oak Quercus robur L.(Fagaceae). Russian Journal of Genetics, 44:997-999.
Richardson, B.J., Baverstock, P.R., Adams, M. (1986). Allozyme electrophoresis: a handbook for animal systematics and population studies. Sydney: Academic Press.
Rousset, F. (1997). Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics, 145:1219-1228.
Slade, D., Skvorc, Z., Ballian, D., Gracan, J., Papes, D. (2008). The Chloroplast DNA Polymorphisms of White Oaks of Section Quercus in The Central Balkans. Silvae Genetica, 57:227.
Stefanova, I., Ammann, B. (2003). Lateglacial and Holocene vegetation belts in the Pirin Mountains (southwestern Bulgaria). The Holocene, 13:97-107.
Stefanović, V., Beus, V., Burlica, Č., Dizdarević, H., Vukorep, I. (1983). Ekološko-vegetacijskarejonizacija Bosnei Hercegovine. Šumarskifakultet u Sarajevu, Posebnaizdanja, 17:1-44.
Streiff, R., Labbe, T., Bacilieri, R., Steinkellner, H., GLÖSSL, J., Kremer, A. (1998). Within‐population genetic structure in Quercus robur L. and Quercus petraea (Matt.) Liebl. assessed with isozymes and microsatellites. Molecular Ecology, 7:317-328.
Tzedakis, P.C. (1994). Vegetation change through glacial—interglacial cycles: a long pollen sequence perspective. Philosophical Transactions of the Royal Society of London. Series B: Biological Sciences, 345:403-432.
Urban, B., Fuchs, M. (2005). Late Pleistocene vegetation of the basin of Phlious, NE-Peloponnese, Greece. Review of Palaeobotany and Palynology, 137:15-29.
Vakkari, P., Blom, A., Rusanen, M., Raisio, J., Toivonen, H. (2006). Genetic variability of fragmented stands of pedunculate oak (Quercus robur) in Finland. Genetica, 127:231-241.
Vakkari, P., Rusanen, M., Heikkinen, J., Huotari, T., Kärkkäinen, K. (2020). Patterns of genetic variation in leading-edge populations of Quercus robur: genetic patchiness due to family clusters. Tree Genetics & Genomes, 16:1-12.
Weir, B.S., Cockerham, A. (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38:1358–1370.
Willis, K.J. (1992). The late Quaternary vegetational history of northwest Greece: I. Lake Gramousti. New Phytologist, 121:101-117.
Yakovlev, I. (2000). Genetic diversity of pedunculate oak (Quercus robur L.) in the middle near Volga region of Russia. Glasnik za Šumske Pokuse, 37:453-468.
Yeh, F.C., Yang, R.C., Boyle, T.B., Ye, Z.H., Mao, J.X. (1997). POPGENE, the user-friendly shareware for population genetic analysis. Molecular biology and biotechnology centre, University of Alberta, Canada, 10:295-301.
Author Info
M. Memiševic Hodžic1, B. Fussi2, D. Gömöry3 and D. Ballian1,4*2Bavarian Office for Forest Genetics (AWG), Teisendorf, Germany
3Technical University in Zvolen, Faculty of Forestry, T. G. Masaryka 24, 96053 Zvolen, Slovakia
4Slovenian Forestry Institute, Vecna pot 2, 1000 Ljubljana, Slovenia
Citation: Hodzic, M.M., Fussi, B., Gomory, D., Ballian, D. (2021). Genetic variability of pedunculate oak (Quercus robur L.) at the Mediterranean margin of the distribution range. Ukrainian Journal of Ecology, 11 (6), 31-40.
Received: 14-Jul-2021 Accepted: 27-Jul-2021 Published: 23-Aug-2021
Copyright: This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
